Single Cell Portal (SCP) is an online portal for sharing single-cell genomics data that is built on top of Terra. Through its searchable database of single-cell studies, interactive visualizations, and sharing tools, SCP supports single-cell researchers throughout the research process. SCP recently published a whitepaper that lays out how the Portal provides this support – this post summarizes the highlights.
Navigating a growing amount of data
Single-cell research is a growing field, with an exponential growth in publications since 2009.
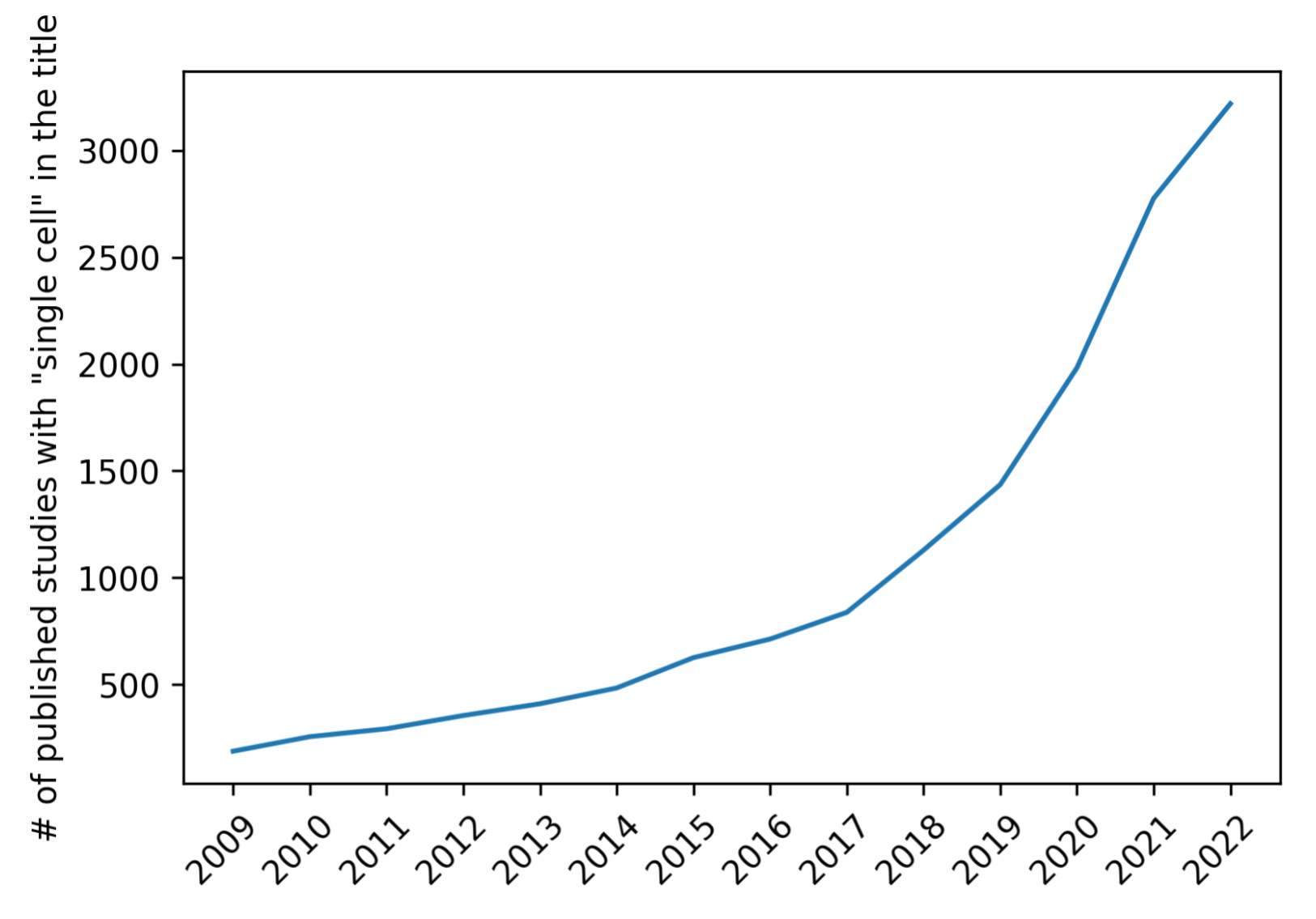

These data are a powerful tool, but they also pose a challenge for researchers seeking to integrate these findings into their own work: it’s hard to sift through hundreds of papers a year to find the data that are relevant. Researchers trying to share their data may also struggle to reach the right audience.
SCP helps researchers navigate these challenges to find and share data more easily.
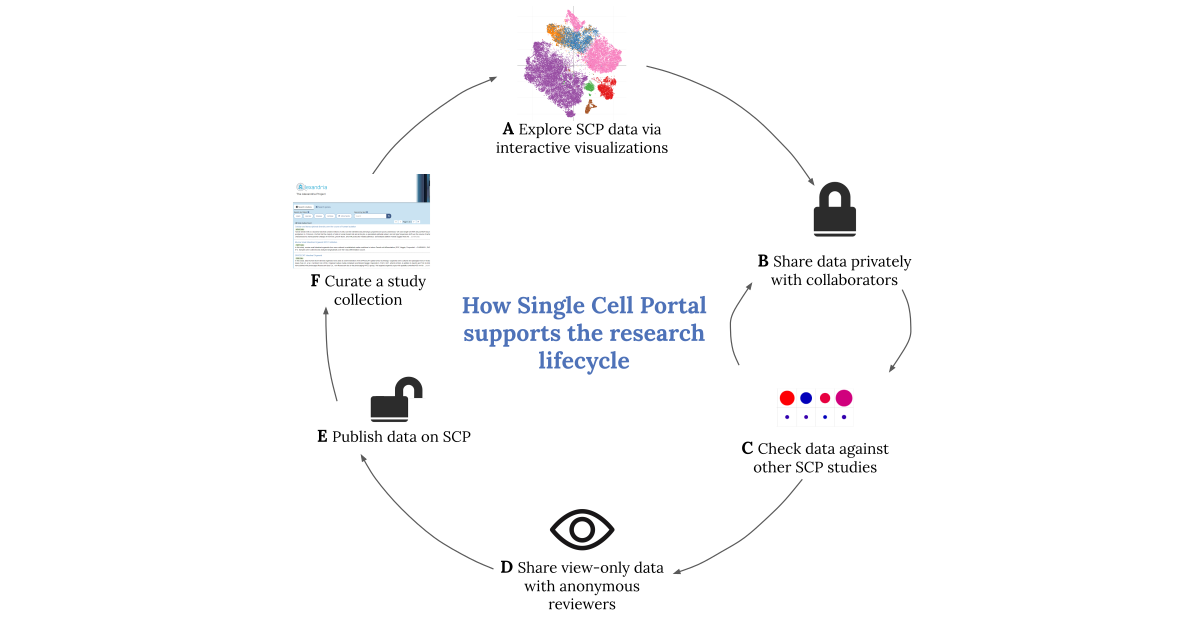

Sparking an idea
SCP’s database provides access to over 500 single-cell studies. Advanced search tools make it easy to zero in on the studies that are relevant to a particular research area. For example, researchers can search by disease, organ, cell type, gene, and library preparation protocol, depending on their questions.
SCP’s interactive visualizations make it easy to explore these datasets, even if researchers don’t have a computational background. This allows users to answer key questions that a static paper figure couldn’t answer, and often more quickly than reading a paper would allow.
Fine-tuning your data
SCP is also a valuable tool for collaborations between computational and non-computational researchers. For example, a computational biologist described how SCP bridges the “wetlab-drylab divide” between collaborators in Single Cell Portal for collaborations: once computationalists upload analysis results to a private SCP study, wet-lab collaborators can explore the data and provide feedback on how to adjust the analysis – without ever having to touch R or a command line.
Sharing results
Once the data are finalized, making them public on SCP increases the work’s impact. An SCP study invites other researchers to engage deeply with the data, exploring it interactively and using it as a reference for patterns observed across studies. Researchers can even download the data – which are hosted within a FISMA-compliant boundary on Terra – and integrate those data into their own analyses.
Try it out!
If you’re intrigued, you can learn more about how Single Cell Portal accelerates single-cell research by reading our whitepaper. You can also visit a demo study to try out the Portal’s interactive visualizations and check out our documentation for details on how to upload your own data to the Portal.



