DRS (pronounced “durs”) stands for Data Repository Service API, one of the interoperability standards developed under the aegis of the Global Alliance for Genomics and Health (GA4GH). Unless you’re an infrastructure nerd, you probably never heard of DRS until you came across something that looked like drs://dg.ANV0/f581ca07-1d7a-4a95-902e-4b910571f3a2 in a data table where you expected something more like gs://some-bucket/blah/some-filename.bam. You may also have encountered Terra documentation that defines DRS URIs, GUIDs, and all that alphabet soup.
What you may not have fully gleaned from those experiences is that DRS is a really neat and phenomenally useful system. It’s also one of those things where you can’t completely avoid having to understand how some cloud stuff works under the hood. So in this blog, we thought it might be helpful to explain the big picture of what DRS is for, and specifically, how it liberates you from having to worry about certain data management problems.
Standardizing data connections across the cloud ecosystem
At its core, DRS is a set of standardized rules for managing how we connect to data in a world that’s increasingly multi-platform and multi-cloud. There are numerous organizations that operate data repositories, often with their own ways of organizing datasets and interfaces for providing access to the data, and hosted on different cloud providers’ platforms, such as Amazon Web Services (AWS), Microsoft Azure, and Google Cloud Platform.
DRS was developed by a GA4GH working group to make it easier for platforms like Terra to connect to those various repositories in order to make the data available for analysis in a way that would be consistent across platforms and sustainable over time. Specifically, one important goal of the standard is to enable individual researchers to access and use data without having to know anything about how the data repositories are engineered, or what are the exact locations of the data files.
That last point, about the location of data files, is what we’re going to dive into next because it’s the only part of DRS that really shows up in a visible way if you’re using Terra, in the form of those somewhat opaque Uniform Resource Identifiers (URIs). The rest all happens behind the scenes, and unless you’re a platform developer or data curator, you don’t need to care about how it works.
Website URLs as a model for understanding how DRS URIs work
The idea here is that DRS acts as an “abstraction layer” that manages the mapping between dataset entries in a database and the actual location of the data files, through the use of URIs. This can be a challenging concept, so let’s take a brief detour to explore what this means through an example that may feel more familiar.
Think about how you interact with a normal website on the internet — like this blog, for example. It’s hosted on a server that has a physical location somewhere, with an IP address that might look something like “172.16.254.1”. But you didn’t have to know that in order to access it. You just typed or clicked on the URL “https://terra.bio/blog/this-article-name”, and here you are. What happened? The URL address you used acted as an alias. Your browser used a domain name service to look up what IP address “terra.bio” corresponds to, and connected you to our server. Then your browser said you wanted this blog article named “this-article-name”, so our server pulled up the relevant content and displayed it in a nice clean webpage, which you’re reading right now.
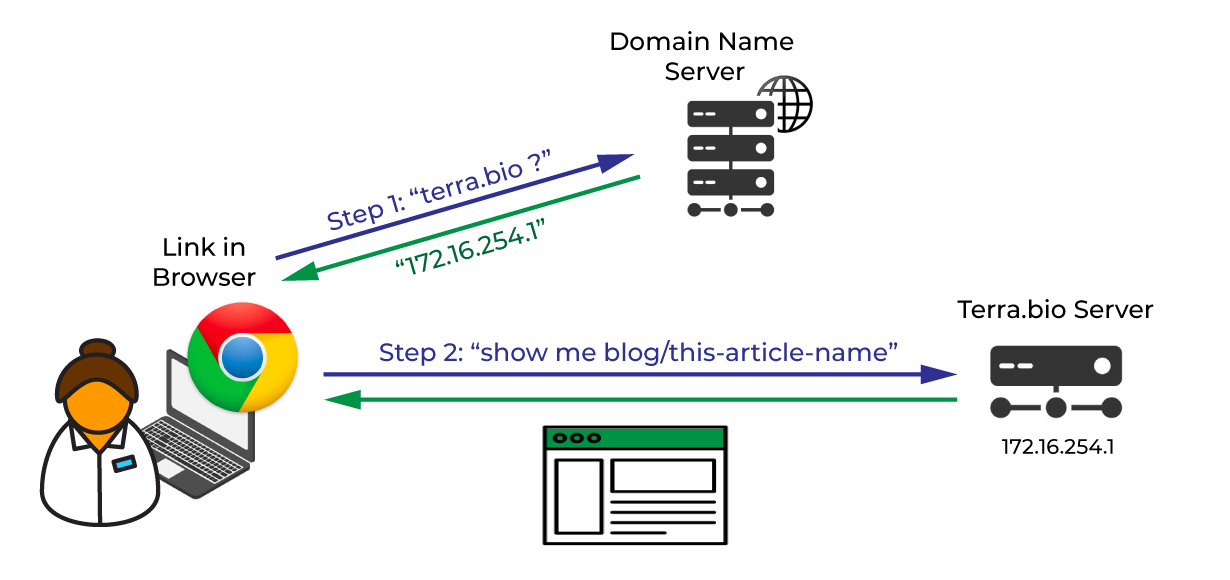

One of the benefits of this system is that, if in six months we need to ditch our current web setup and move to a different server or service provider, we can do that behind the scenes. We’d just update the location that the terra.bio alias should point to in the domain name service list, and you would never even need to know. Technically that’s exactly what we did in December when we launched the new Terra.bio website: we moved from Squarespace to WP Engine, and anyone who had bookmarked Terra.bio got automatically directed to the new location without having to do anything. A similar mechanism applies to content on the server as well: if we decide to change how we manage our content (whether it’s how files are named, where they live, or how the database is organized) we can set up aliases so that any existing links will continue to work seamlessly.
And there you have it: that’s essentially what DRS does for us in the context of accessing data in various repositories. Repositories that use DRS for managing data connections assign a sort of alias called a Uniform Resource Identifier (URI) to each data file so that a platform like Terra can provide access to the data in a way that is agnostic of the actual location of the files.
Using data from repositories with and without DRS
Let’s run through an example. Say you want to run an analysis on a dataset from a cloud repository included in the Terra catalog. When you import the data into your Terra workspace, the system creates one or more data tables that act as a manifest containing all the relevant metadata, including references to any data files.
What it looks like without DRS
For a “regular” repository that does not use DRS, the locations in the table are the actual full paths to where the files are stored. Each file location starts with the appropriate prefix for the storage system (gs:// for Google Cloud Storage), the name of the storage bucket (fc-***/), and the full path to the file (everything else).
gs://fc-56ac46ea-efc4-4683-b6d5-6d95bed41c5e/CCDG_13607/Project_CCDG_13607_B01_GRM_WGS.cram.2019-02-06/Sample_NA20770/analysis/NA20770.final.cram(Yeah it’s a lot.)
When you launch your analysis on the data, the system uses those file locations to access the relevant files. So you run a workflow, get results, move on with your life, yay.
Then three months later you discover you used the wrong parameter in your pipeline and you go to run the workflow again, but it fails. The error message says the files could not be found. After some digging, you find out that the file locations have changed because the data custodians decided to reorganize their data content, or changed the naming conventions of the files, or whatever. As a consequence, you have to look up the new locations for the data you’re interested in and update your data tables. It’s a huge hassle.
DRS to the rescue
Now imagine you live in a parallel universe where that same repository uses DRS to manage data location. You get the same data tables after importing the data into your workspace, but this time the file location column contains DRS URIs instead of the actual file locations. Each starts with the DRS prefix (drs://), an identifier for the repository (dg.ANV0/), and an identifier specific to the file itself (everything else).
drs://dg.ANV0/f581ca07-1d7a-4a95-902e-4b910571f3a2(Not exactly human-readable, but at least it’s shorter than the full path!)
When you encounter these for the first time it can be quite confusing, but good news: you can plug those URIs straight into your workflow configuration where you would normally put the file locations. When you launch your workflow, Terra’s workflow management system will recognize that these are DRS URIs because of the “drs://” prefix. Terra will then identify and connect to the original repository, where it will look up each file’s URI and get the actual file location associated with that URI. Then the rest of the work can proceed as normal — get results, move on, yay.
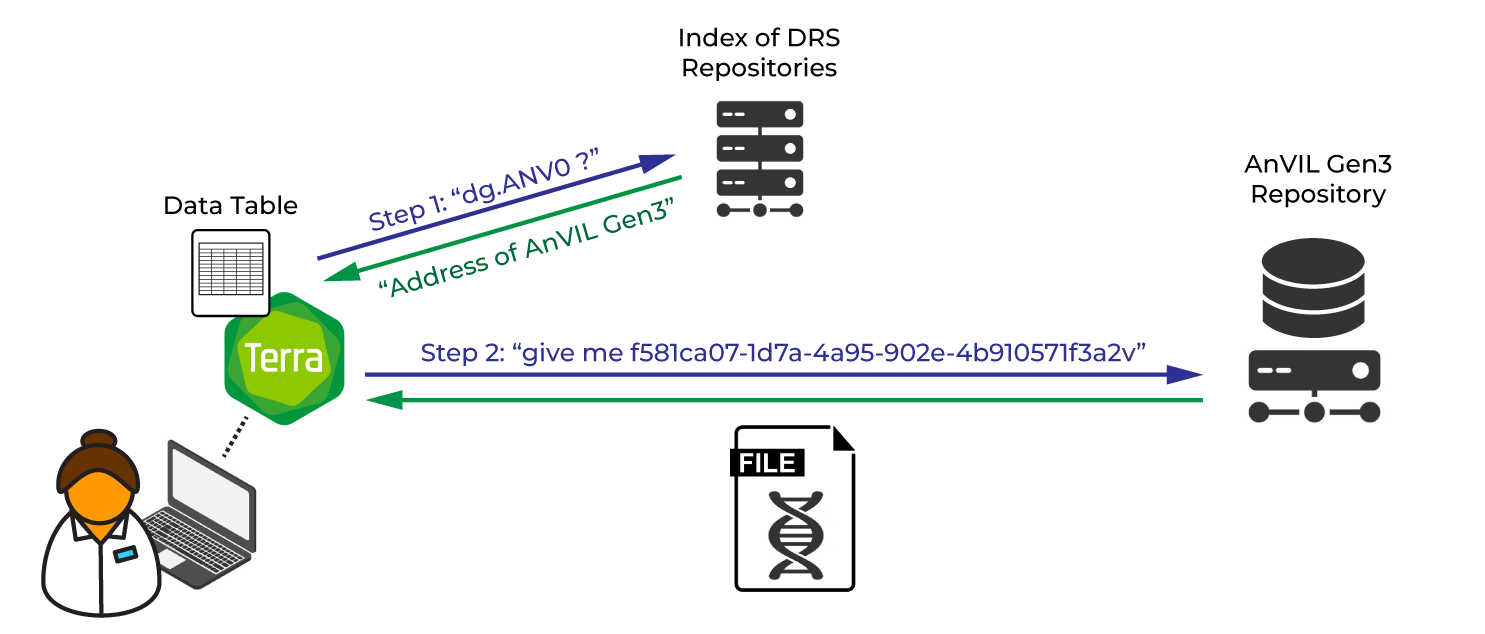

And hey, three months later, when you re-run the analysis with the corrected parameters, everything still works fine and you don’t even realize the location of the data has changed. The data custodians updated the DRS manifests for their dataset with the new locations, so the original URIs still work. Thanks, DRS.
Note that if you’re using the data in a Notebook or in Rstudio, you have to invoke some code to transform the DRS URIs into file locations; it’s not as completely handled for you as with workflows. But there’s a handy package you can call on, so it’s reasonably straightforward, and it’s all documented here.
A win for interoperability and multi-cloud support
Hopefully, by now I’ve convinced you that DRS is a really useful system and that it’s worth the effort of learning to work with its URIs, if only because it helps ensure that references to data files will remain valid throughout the lifetime of your analysis.
On top of that, it’s also going to be very helpful as we move closer to multi-cloud support. Datasets of widespread biomedical importance are increasingly made available on more than one cloud platform, in order to ensure access regardless of what cloud provider a researcher may be tied to. Thanks to DRS, data custodians don’t have to maintain separate data manifests for the mirrored datasets, because DRS URIs can be associated with multiple file locations corresponding to different platforms. As a result, a DRS-enabled repository can route incoming URI-based requests for data to the appropriate file locations depending on what platform the requester is calling from. The benefit for you as a researcher is that you’ll be able to use the same exact data tables to run workflows on either GCP or Azure, once Azure support comes online in Terra.
The last mile of the DRS learning curve
You now know all the major points of what DRS entails and how it shows up in Terra. You can learn more about using DRS URIs in practice from this Terra documentation article, and see them in action in several public workspaces: the Terra-201-Gen3-Module workspace provides instructions for retrieving data from a DRS-enabled repository (AnVIL Gen3), and the BioData Catalyst Collection workspace features several tables that reference files using DRS URIs (look for a column named “pfb:ga4gh_drs_uri”). Note that the repositories currently using DRS in the Terra catalog all require authentication, so you’ll need to go through a simple login step to “link” your Terra credentials with the corresponding repository system. This is described at the very end of the documentation article, in the section titled “Troubleshooting DRS URI access in Terra”.
As always, if you run into any trouble, don’t hesitate to reach out to the Terra Helpdesk!
Additional resources
DRS API: Enabling Cloud-Based Data Access and Retrieval
GA4GH 2020 Connection Demos Highlight the Value of Interoperability in Genomics



